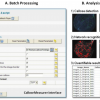
Callose Measurer
Robatzek lab @ Sainsbury Laboratory
Overview
| Measured variables | callose-deposition |
|---|---|
| Operating system | windows |
| Licence | freeware |
| Automation level | automated |
| Plant requirements | any |
| Export formats | unknown |
| Other information | Requires Acapella software |
Scientific article(s)
CalloseMeasurer: a novel software solution to measure callose deposition and recognise spreading callose patternsJi Zhou,Thomas Spallek,Christine Faulkner,Silke RobatzekPlant Methods, 2012 View paper
Similar tools
Other tools for the analysis of cell:
Description
CalloseMeasurer is a new software solution for batch-processing large image data sets to quantify callose deposition in plants, a plant immune response triggered by potentially pathogenic microbes. Additionally, by tracking identified callose deposits between multiple images, the software can recognise patterns of how a given filamentous pathogen grows in plant leaves. The software has been evaluated with typical noisy experimental images and can be automatically executed without the need for user intervention. The automated analysis is achieved by using standard image analysis functions such as image enhancement, adaptive thresholding, and object segmentation, supplemented by several novel methods which filter background noise, split fused signals, perform edge-based detection, and construct networks and skeletons for extracting pathogen growth patterns. To efficiently batch process callose images, we implemented the algorithm in C/C++ within the Acapella™ framework. Using the tool we can robustly score significant differences between different plant genotypes when activating the immune response. We also provide examples for measuring the in planta hyphal growth of filamentous pathogens. The software is an easy-to-use protocol which is executed within the Acapella software system without requiring any additional libraries.
Source: Zhou et al. (2012) Plant methods , 8(1), 49